Clinical Significance of 21q22.2 Genomic Rearrangement in Prostate Cancer
Authors: Francesco Giuseppe Carbone1, Mattia Barbareschi1 and Andrea Lunardi2,*
1Unit of Surgical Pathology, Santa Chiara Hospital, Trento, Italy
2Department of Cellular, Computational and Integrative Biology-CIBIO, University of Trento, Trento, Italy.
*Correspondence to: Professor. Andrea Lunardi, Department of Cellular, Computational and Integrative Biology-CIBIO, University of Trento, Via Sommarive 9, 38123 Trento, Italy; E-mail: andrea.lunardi@unitn.it
Received: 20 October 2022; Accepted: 21 November 2022; Published: 28 November 2022
ORCID
Francesco Carbone: https://orcid.org/0000-0001-8425-9658
Mattia Barbareschi: https://orcid.org/0000-0003-0217-3223
Andrea Lunardi: https://orcid.org/0000-0001-6218-2565
Citation: Carbone FG, Barbareschi M, Lunardi A (2022) Clinical Significance of 21q22.2 Genomic Rearrangement in Prostate Cancer, 21st Century Pathology, Volume 2 (6): 132
Abstract
Prostate cancer (PCa) is a heterogeneous disease characterized by highly recurrent molecular alterations. Among them, the 21q22-23 intrachromosomal rearrangement leading to TMPRSS2-ERG gene fusion is unquestionably the most common. De novo expression of the N-terminal truncated form of ERG (ERGMet40) provides PCa cells with aggressiveness and metastatic potential. Pre-clinical evidence defines ERG expression as a vulnerable condition for synthetic lethality approaches. However, clinical trials failed in demonstrating greater efficacy of PARP inhibitors in ERG tumors, leaving many questions unanswered. Organoid technology and artificial intelligence may help find the solutions.
Keywords:
Prostate Cancer; ERG; Organoids; Synthetic Lethality
Abbreviations: PCa: Prostate Cancer; mCRPC: metastatic Castration Resistant Prostate Cancer; HGPIN: High-Grade Intraepithelial Neoplasia; AIP: atypical intraductal proliferations; IDC: Intraductal carcinoma; PrOs: Prostate Organoids.
Introduction
Occurring in more than 50% of cases, rearrangements of members of the ETS-gene family are the most common genetic alterations in prostate cancer (PCa) [1]. Among them, the predominant is the 21q22.2-3 that involves the Transmembrane Protease Serine 2 (TMPRSS2) and ETS-related gene (ERG). Detectable in almost 50% of prostate cancer biopsies from Caucasians, TMPRRS2-ERG genes fusion is less frequent in African-American and Asian men (27-31%) [2]. Moreover, the rearrangement is commonly identified in 5-30% of high-grade PIN (HGPIN) lesions, which classifies the lesion as an early event during prostate carcinogenesis [3-5]. TMPRSS2-ERG rearrangement characterizes as well other prostatic histotypes -such as small cell carcinoma of the prostate (pure or mixed with acinar carcinoma)- where it’s found in about 45% of cases and where its presence underscores the prostatic origin of the malignancy in the differential diagnosis of a metastatic small cell carcinoma of unknown primary origin [6,7]. Functionally, ERG expression has been associated to tumor aggressiveness by promoting local invasion and metastatic progression of the disease through the transcriptional control of targeted genes in PCa cells [8-10]. Less clear is the role of ERG in the early stage of prostate tumorigenesis since in vitro and in vivo studies demonstrate that ERG is not sufficient to trigger the malignant transformation of the prostate epithelium [9-14].
Leveraging organoid technology to investigate ERG functions
Prostate organoids (PrOs) are cell culture models constituted by proliferating wild type adult prostate progenitors that self-assemble to reconstitute the correct basal-luminal architecture surrounding the lumen of adult prostate epithelium. Notably, PrOs are characterized by a very stable genome and dependence on testosterone for luminal lineage differentiation and lumenogenesis [15]. Doxycycline inducible expression of exogenous ERGMet40 in mouse prostate organoids (mPrOs) determines the expansion of the luminal compartment and contraction of the basal layer, closely resembling the histologic appearance of HGPIN. However, ERG+ organoids grow slowly compared to control lines and they are likely to get lost after a few passages if kept in full medium. Surprisingly, in sharp contrast to wild-type prostate organoids that are dependent on exogenous epidermal growth factor (EGF, 0.5 ng/mL) to survive, ERG+mPrOs grow in the absence of EGF [16], supporting the thesis that ERG+HGPIN cells could escape the quiescence that largely characterizes adult prostate tissue homeostasis and proliferate in the absence of physiologic stimuli. HGPIN is not the only intraductal proliferative lesion of the prostate. Intraductal carcinoma (IDC) is a clinically aggressive form of prostatic carcinoma characterized by a lumen-spanning proliferation of neoplastic prostate epithelium
within a preserved basal layer and it is associated with a high Gleason score (>7), large tumor volume, extra prostatic extension, positive lymph node status, and increased recurrence risk [17-19]. Most (about 75%) IDC lesions have the TMPRSS2-ERG rearrangement [20] and a similar percentage is found in atypical intraductal cribriform proliferations (AIP) that histologically appear more ominous than HGPIN, but do not fulfil the criteria of classic IDC [21] (Figure 1).
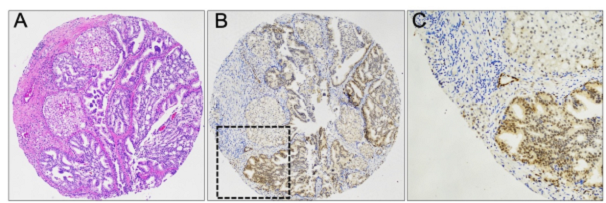
Figure 1: Atypical intraductal proliferation (AIP) intermingled with intraductal carcinoma (IDC), both with dense cribriform pattern (A, H&E x10). Low (B, x10) and high (C, x40) magnification of ERG immunohistochemistry showing diffuse nuclear positivity.
Recent studies have shown that prostate cancer with coexistent IDC and/or invasive cribriform carcinoma is associated with a higher percentage of genomic alterations than prostate cancer without these patterns [22,23], and that such genomic events cluster in specific chromosomal regions associated with aggressive disease such as deletions of 8p, with the involvement of the NKX3.1 gene [22]. In addition to its role as a transcription factor, NKX3.1 protein contributes to genome stability by favoring DNA damage repair in prostate epithelium [24-28] while ERG is known to promotes DNA double strand breaks (DSBs) [29-31]. In prostate progenitors, we demonstrated that ERG pose a major threat to genomic stability by promoting the proteasome degradation of NKX3.1 protein and the consequent accumulation of ERG-induced DNA damage, which remains sub-lethal [16]. These findings could reasonably help explain, at least in part, the increased rate of genomic alterations and highly malignant behavior of IDC and invasive cribriform carcinoma characterized by TMPRSS2-ERG rearrangement and NKX3.1 loss.
Clinical relevance of ETS genes expression
As ERG expression in prostate cancer cells leads to DNA DSB [30,31], synthetic lethality strategies of therapeutic intervention should be considered in patients with tumors harboring ETS related gene rearrangement [32]. sTOPARP-B trial showed anti-tumor activity of the PARP-inhibitor olaparib in patients with specific DNA repair gene defects (DRDs) [33]. Similar conclusions have been obtained by the PROFOUND trial for patients having somatic or germinal DRDs [34,35]. GALAHAD (a phase II trial) demonstrated a significant response rate of mCRPC patient with BRCA mutations treated with niraparib [36]. Finally, again in patients presenting with DRDs, promising results have been obtained by TRITON2 and TALAPRO-1 trials with the use of rucaparib and talazoparib, respectively [37-39]. As shown by Brenner JC, et al. (2011) treatment of ERG overexpressing cells with the PARP inhibitor olaparib not only decreased ERG-mediated cell invasion and intravasation, but also inhibited growth in mouse xenograft models [30]. Further in vitro work showed resistance to radiation of cells overexpressing ERG, which was reverted by inhibition of PARP [31,40]. However, clinical trials failed to identify differences in the response rate of ETS (mainly ERG) positive and negative mCRPC treated with PARP inhibitor veliparib and the androgen biosynthesis inhibitor abiraterone [41]. Similarly, ERG fusion status was not prognostic in patients with intermediate risk prostate cancer treated with radiation [42], although the presence of ERG rearrangement and PTEN loss was independently associated with recurrence-free survival in patients undergoing brachytherapy [43]. Future studies might clarify the potential role of ERG in selecting patients who could benefit more from a PARP inhibitor therapy, alone or in combination with other therapies. Along with PARPi, in the last few years immunotherapy has become increasingly important in the therapeutic plan of oncological patients, in particular the use of immune checkpoint-inhibitors (ICIs). Programmed death-1 (PD-1) and its ligand PD-L1 are transmembrane glycoproteins expressed by different types of immune cells, and, mostly PD-L1, by tumor cells including those of prostate cancer [44]. The PD-1/PD-L1 pathway leads to the inactivation of PD-1 expressing cells, mainly CD8+ cytotoxic T cells, thus favoring tumor immune escape [44]. Accordingly, increasing attention has been dedicated to immunohistochemical evaluation of PD-1/PD-L1 proteins as predictor factors of ICIs efficacy. Therapeutic strategy based on ICIs proved to be effective for various cancer types including non-small-cell lung cancer [45], renal cell carcinoma [46,47], urothelial cancer [48,49], colorectal cancer [50], breast cancer [51,52] and melanoma [53,54], upon accurate stratification of patients. Contrariwise, ICIs resulted poorly effective in prostate cancer patients, likely dependent by the strong immunosuppressive tumor microenvironment (TME), the lower infiltration of T-cells and the reduced tumor mutation burden (TMB) [55]. Recent clinical trials, however, have demonstrated excellent responses to ICIs and/or their combinations with other agents of prostate cancer characterized by the biallelic loss of CDK12, high tumor mutation burden, high microsatellite instability (MSI-h) and mismatch repair-deficient (dMMR) [55,56]. ERG expression can also contribute to increase genome instability and tumor mutation burden in PCa cells [16]. In this scenario, it is interesting to note that a concordant IHC status of ERG with a “nodular” pattern of PDL1 has been described [44,57].
Further studies are needed to investigate the role of ERG fusions with regards to PD-L1/PD-1 expression, immune infiltration, and ICIs response.
Conclusion
One critical aspect may be the criteria adopted for ERG stratification. Rather than “positive” or “negative”, a quantitative biparametric assessment by IHC of the different levels of ETS protein expression and the relative percentage of positive cells might help to better interpret the response of ETS-positive tumors to specific clinical protocol [58]. Artificial intelligence is fueling the field of computational quantitative pathology, which will contribute substantially to solving issues like this. Machine learning-based approaches will improve the accuracy and reproducibility of histopathological analyses, while also making them faster and, overall, quantitative [59]. Further clinical studies will be needed to understand the complex relationship between the expression of ETS genes and the clinical relevance of agents acting on the DNA damage response and immune activity. In the meantime, the goal of preclinical research will be to increasingly define the molecular mechanisms that undermine susceptibility to PARP and IC inhibitors, with the aim of discovering new therapeutic strategies and expanding the potential spectrum of treatable patients (Figure 2).
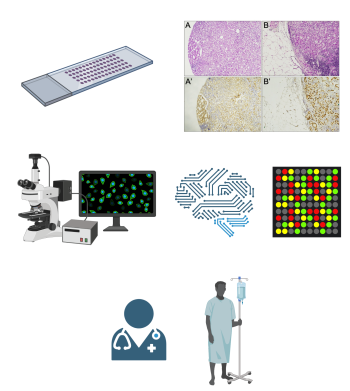
Figure 2: Prostate carcinoma (A, H&E x20) and lymph node metastasis (B, H&E x20) positive for ERG (A', B'; IHC x20). Note that ERG shows different degrees of expression within the same specimen and between distinct specimens (created with BioRender.com).
Author Contributions
Conceptualization: AL, FGC. MB. Data curation: FGC. Formal analysis: FGC, MB, AL. Funding acquisition: AL. Visualization: FGC, AL. Writing – original draft: FGC. Writing – review & editing: AL, FGC, MB.
Conflict of Interest
The authors declare no conflicts of interest.
References
1. Abeshouse A, Ahn J, Akbani R, Ally A, Amin S, Andry CD, Annala M, Aprikian A, Armenia J, Arora A, Auman JT. The molecular taxonomy of primary prostate cancer. Cell. 2015 Nov 5;163(4):1011-25. https://doi.org/10.1016/j.cell.2015.10.025
2. Magi?Galluzzi C, Tsusuki T, Elson P, Simmerman K, LaFargue C, Esgueva R, Klein E, Rubin MA, Zhou M. TMPRSS2–ERG gene fusion prevalence and class are significantly different in prostate cancer of caucasian, african?american and japanese patients. The Prostate. 2011 Apr;71(5):489-97. https://doi.org/10.1002/pros.21265
3. Perner S, Mosquera JM, Demichelis F, Hofer MD, Paris PL, Simko J, Collins C, Bismar TA, Chinnaiyan AM, De Marzo AM, Rubin MA. TMPRSS2-ERG fusion prostate cancer: an early molecular event associated with invasion. The American journal of surgical pathology. 2007 Jun 1;31(6):882-8. https://doi.org/10.1097/01.pas.0000213424.38503.aa
4. Morais CL, Guedes LB, Hicks J, Baras AS, De Marzo AM, Lotan TL. ERG and PTEN status of isolated high-grade PIN occurring in cystoprostatectomy specimens without invasive prostatic adenocarcinoma. Human pathology. 2016 Sep 1;55:117-25. https://doi.org/10.1016/j.humpath.2016.04.017
5. Mosquera JM, Perner S, Genega EM, Sanda M, Hofer MD, Mertz KD, Paris PL, Simko J, Bismar TA, Ayala G, Shah RB. Characterization of TMPRSS2-ERG fusion high-grade prostatic intraepithelial neoplasia and potential clinical implications. Clinical Cancer Research. 2008 Jun 1;14(11):3380-5. https://doi.org/10.1158/1078-0432.CCR-07-5194
6. Lotan TL, Gupta NS, Wang W, Toubaji A, Haffner MC, Chaux A, Hicks JL, Meeker AK, Bieberich CJ, De Marzo AM, Epstein JI. ERG gene rearrangements are common in prostatic small cell carcinomas. Modern pathology. 2011 Jun;24(6):820-8. https://doi.org/10.1038/modpathol.2011.7
7. Scheble VJ, Braun M, Wilbertz T, Stiedl AC, Petersen K, Schilling D, Reischl M, Seitz G, Fend F, Kristiansen G, Perner S. ERG rearrangement in small cell prostatic and lung cancer. Histopathology. 2010 Jun;56(7):937-43. https://doi.org/10.1111/j.1365-2559.2010.03564.x
8. Demichelis F, Rubin MA. TMPRSS2-ETS fusion prostate cancer: biological and clinical implications. Journal of clinical pathology. 2007 Nov 1;60(11):1185-6. http://dx.doi.org/10.1136/jcp.2007.046557
9. Carver BS, Tran J, Gopalan A, Chen Z, Shaikh S, Carracedo A, Alimonti A, Nardella C, Varmeh S, Scardino PT, Cordon-Cardo C. Aberrant ERG expression cooperates with loss of PTEN to promote cancer progression in the prostate. Nature genetics. 2009 May;41(5):619-24. https://doi.org/10.1038/ng.370
10. Alaimo A, Lorenzoni M, Ambrosino P, Bertossi A, Bisio A, Macchia A, Zoni E, Genovesi S, Cambuli F, Foletto V, De Felice D. Calcium cytotoxicity sensitizes prostate cancer cells to standard-of-care treatments for locally advanced tumors. Cell death & disease. 2020 Dec 7;11(12):1-7. https://doi.org/10.1038/s41419-020-03256-5
11. Klezovitch O, Risk M, Coleman I, Lucas JM, Null M, True LD, Nelson PS, Vasioukhin V. A causal role for ERG in neoplastic transformation of prostate epithelium. Proceedings of the National Academy of Sciences. 2008 Feb 12;105(6):2105-10. https://doi.org/10.1073/pnas.0711711105
12. King JC, Xu J, Wongvipat J, Hieronymus H, Carver BS, Leung DH, Taylor BS, Sander C, Cardiff RD, Couto SS, Gerald WL. Cooperativity of TMPRSS2-ERG with PI3-kinase pathway activation in prostate oncogenesis. Nature genetics. 2009 May;41(5):524-6. https://doi.org/10.1038/ng.371
13. Chen Y, Chi P, Rockowitz S, Iaquinta PJ, Shamu T, Shukla S, Gao D, Sirota I, Carver BS, Wongvipat J, Scher HI. ETS factors reprogram the androgen receptor cistrome and prime prostate tumorigenesis in response to PTEN loss. Nature medicine. 2013 Aug;19(8):1023-9. https://doi.org/10.1038/nm.3216
14. Tomlins SA, Laxman B, Varambally S, Cao X, Yu J, Helgeson BE, Cao Q, Prensner JR, Rubin MA, Shah RB, Mehra R. Role of the TMPRSS2-ERG gene fusion in prostate cancer. Neoplasia. 2008 Feb 1;10(2):177-IN9. https://doi.org/10.1593/neo.07822
15. Cambuli F, Foletto V, Alaimo A, De Felice D, Gandolfi F, Palumbieri MD, Zaffagni M, Genovesi S, Lorenzoni M, Celotti M, Bertossio E. Intra?epithelial non?canonical Activin A signaling safeguards prostate progenitor quiescence. EMBO reports. 2022 May 4;23(5):e54049. https://doi.org/10.15252/embr.202154049
16. Lorenzoni M, De Felice D, Beccaceci G, Di Donato G, Foletto V, Genovesi S, Bertossi A, Cambuli F, Lorenzin F, Savino A, Avalle L. ETS-related gene (ERG) undermines genome stability in mouse prostate progenitors via Gsk3β dependent Nkx3. 1 degradation. Cancer letters. 2022 May 28;534:215612. https://doi.org/10.1016/j.canlet.2022.215612
17. Miyai K, Divatia MK, Shen SS, Miles BJ, Ayala AG, Ro JY. Clinicopathological analysis of intraductal proliferative lesions of prostate: intraductal carcinoma of prostate, high-grade prostatic intraepithelial neoplasia, and atypical cribriform lesion. Human pathology. 2014 Aug 1;45(8):1572-81. https://doi.org/10.1016/j.humpath.2014.03.011
18. Van der Kwast T, Al Daoud N, Collette L, Sykes J, Thoms J, Milosevic M, Bristow RG, Van Tienhoven G, Warde P, Mirimanoff RO, Bolla M. Biopsy diagnosis of intraductal carcinoma is prognostic in intermediate and high risk prostate cancer patients treated by radiotherapy. European journal of cancer. 2012 Jun 1;48(9):1318-25. https://doi.org/10.1016/j.ejca.2012.02.003
19. Wilcox G, Soh S, Chakraborty S, Scardino PT, Wheeler TM. Patterns of high-grade prostatic intraepithelial neoplasia associated with clinically aggressive prostate cancer. Human pathology. 1998 Oct 1;29(10):1119-23. https://doi.org/10.1016/S0046-8177(98)90423-3
20. Han B, Suleman K, Wang L, Siddiqui J, Sercia L, Magi-Galluzzi C, Palanisamy N, Chinnaiyan AM, Zhou M, Shah RB. ETS gene aberrations in atypical cribriform lesions of the prostate: implications for the distinction between intraductal carcinoma of the prostate and cribriform high-grade prostatic intraepithelial neoplasia. The American journal of surgical pathology. 2010 Apr 1;34(4):478-85. https://doi.org/10.1097/PAS.0b013e3181d6827b
21. Hickman RA, Yu H, Li J, Kong M, Shah RB, Zhou M, Melamed J, Deng FM. Atypical intraductal cribriform proliferations of the prostate exhibit similar molecular and clinicopathologic characteristics as intraductal carcinoma of the prostate. The American Journal of Surgical Pathology. 2017 Apr 1;41(4):550-6. https://doi.org/10.1097/PAS.0000000000000794
22. Böttcher R, Kweldam CF, Livingstone J, Lalonde E, Yamaguchi TN, Huang V, Yousif F, Fraser M, Bristow RG, van der Kwast T, Boutros PC. Cribriform and intraductal prostate cancer are associated with increased genomic instability and distinct genomic alterations. BMC cancer. 2018 Dec;18(1):1-1. https://doi.org/10.1186/s12885-017-3976-z
23. Chua ML, Lo W, Pintilie M, Murgic J, Lalonde E, Bhandari V, Mahamud O, Gopalan A, Kweldam CF, van Leenders GJ, Verhoef EI. A prostate cancer “nimbosus”: genomic instability and SChLAP1 dysregulation underpin aggression of intraductal and cribriform subpathologies. European urology. 2017 Nov 1;72(5):665-74. https://doi.org/10.1016/j.eururo.2017.04.034
24. Papachristodoulou A, Rodriguez-Calero A, Panja S, Margolskee E, Virk RK, Milner TA, Martina LP, Kim JY, Di Bernardo M, Williams AB, Maliza EA. NKX3. 1 localization to mitochondria suppresses prostate cancer initiation. Cancer discovery. 2021 Sep;11(9):2316-33. https://doi.org/10.1158/2159-8290.CD-20-1765
25. Zhang H, Zheng T, Chua CW, Shen M, Gelmann EP. Nkx3. 1 controls the DNA repair response in the mouse prostate. The Prostate. 2016 Mar;76(4):402-8. https://doi.org/10.1002/pros.23131
26. Bowen C, Gelmann EP. NKX3. 1 activates cellular response to DNA damage. Cancer research. 2010 Apr 15;70(8):3089-97. https://doi.org/10.1158/0008-5472.CAN-09-3138
27. Bowen C, Ju JH, Lee JH, Paull TT, Gelmann EP. Functional activation of ATM by the prostate cancer suppressor NKX3. 1. Cell reports. 2013 Aug 15;4(3):516-29. https://doi.org/10.1016/j.celrep.2013.06.039
28. Ouyang X, DeWeese TL, Nelson WG, Abate-Shen C. Loss-of-function of Nkx3. 1 promotes increased oxidative damage in prostate carcinogenesis. Cancer research. 2005 Aug 1;65(15):6773-9. https://doi.org/10.1158/0008-5472.CAN-05-1948
29. Chatterjee P, Choudhary GS, Alswillah T, Xiong X, Heston WD, Magi-Galluzzi C, Zhang J, Klein EA, Almasan A. The TMPRSS2–ERG Gene Fusion Blocks XRCC4-Mediated Nonhomologous End-Joining Repair and Radiosensitizes Prostate Cancer Cells to PARP InhibitionTMPRSS2–ERG Fusion Inhibits NHEJ DNA Repair. Molecular cancer therapeutics. 2015 Aug 1;14(8):1896-906. https://doi.org/10.1158/1535-7163.MCT-14-0865
30. áChad Brenner J, Ateeq B, Li Y, Yocum A, Cao Q, Asangani I, Patel S, Wang X, Liang H, Yu J, Palanisamy N. Mechanistic rationale for inhibition of poly (ADP-ribose) polymerase in ETS gene fusion-positive prostate cancer. Cancer cell. 2011 May 17;19(5):664-78. https://doi.org/10.1016/j.ccr.2011.04.010
31. Chatterjee P, Choudhary GS, Sharma A, Singh K, Heston WD, Ciezki J, Klein EA, Almasan A. PARP inhibition sensitizes to low dose-rate radiation TMPRSS2-ERG fusion gene-expressing and PTEN-deficient prostate cancer cells. PloS one. 2013 Apr 2;8(4):e60408. https://doi.org/10.1371/journal.pone.0060408
32. Lorenzin F, Demichelis F. Past, Current, and Future Strategies to Target ERG Fusion-Positive Prostate Cancer. Cancers. 2022 Feb 22;14(5):1118. https://doi.org/10.3390/cancers14051118
33. Mateo J, Porta N, Bianchini D, McGovern U, Elliott T, Jones R, Syndikus I, Ralph C, Jain S, Varughese M, Parikh O. Olaparib in patients with metastatic castration-resistant prostate cancer with DNA repair gene aberrations (TOPARP-B): a multicentre, open-label, randomised, phase 2 trial. The lancet oncology. 2020 Jan 1;21(1):162-74. https://doi.org/10.1016/S1470-2045(19)30684-9
34. Hussain M, Mateo J, Fizazi K, Saad F, Shore N, Sandhu S, Chi KN, Sartor O, Agarwal N, Olmos D, Thiery-Vuillemin A. Survival with olaparib in metastatic castration-resistant prostate cancer. New England Journal of Medicine. 2020 Dec 10;383(24):2345-57. https://doi.org/10.1056/NEJMoa2022485
35. de Bono J, Mateo J, Fizazi K, Saad F, Shore N, Sandhu S, Chi KN, Sartor O, Agarwal N, Olmos D, Thiery-Vuillemin A. Olaparib for metastatic castration-resistant prostate cancer. New England Journal of Medicine. 2020 May 28;382(22):2091-102. https://doi.org/10.1056/NEJMoa1911440
36. Smith MR, Scher HI, Sandhu S, Efstathiou E, Lara Jr PN, Evan YY, George DJ, Chi KN, Saad F, Ståhl O, Olmos D. Niraparib in patients with metastatic castration-resistant prostate cancer and DNA repair gene defects (GALAHAD): a multicentre, open-label, phase 2 trial. The Lancet Oncology. 2022 Mar 1;23(3):362-73. https://doi.org/10.1016/S1470-2045(21)00757-9
37. Abida W, Patnaik A, Campbell D, Shapiro J, Bryce AH, McDermott R, Sautois B, Vogelzang NJ, Bambury RM, Voog E, Zhang J. Rucaparib in men with metastatic castration-resistant prostate cancer harboring a BRCA1 or BRCA2 gene alteration. Journal of Clinical Oncology. 2020 Nov 11;38(32):3763. https://doi.org/10.1200/JCO.20.01035
38. Abida W, Campbell D, Patnaik A, Shapiro JD, Sautois B, Vogelzang NJ, Voog EG, Bryce AH, McDermott R, Ricci F, Rowe J. Non-BRCA DNA Damage Repair Gene Alterations and Response to the PARP Inhibitor Rucaparib in Metastatic Castration-Resistant Prostate Cancer: Analysis From the Phase II TRITON2 StudyRucaparib in mCRPC with a Non-BRCA DDR Gene Alteration. Clinical Cancer Research. 2020 Jun 1;26(11):2487-96. https://doi.org/10.1158/1078-0432.CCR-20-0394
39. de Bono JS, Mehra N, Scagliotti GV, Castro E, Dorff T, Stirling A, Stenzl A, Fleming MT, Higano CS, Saad F, Buttigliero C. Talazoparib monotherapy in metastatic castration-resistant prostate cancer with DNA repair alterations (TALAPRO-1): An open-label, phase 2 trial. The Lancet Oncology. 2021 Sep 1;22(9):1250-64. https://doi.org/10.1016/S1470-2045(21)00376-4
40. Han S, Brenner JC, Sabolch A, Jackson W, Speers C, Wilder-Romans K, Knudsen KE, Lawrence TS, Chinnaiyan AM, Feng FY. Targeted radiosensitization of ETS fusion-positive prostate cancer through PARP1 inhibition. Neoplasia. 2013 Oct 1;15(10):1207-IN36. https://doi.org/10.1593/neo.131604
41. Hussain M, Daignault-Newton S, Twardowski PW, Albany C, Stein MN, Kunju LP, Siddiqui J, Wu YM, Robinson D, Lonigro RJ, Cao X. Targeting androgen receptor and DNA repair in metastatic castration-resistant prostate cancer: results from NCI 9012. Journal of Clinical Oncology. 2018 Apr 4;36(10):991. https://doi.org/10.1200/JCO.2017.75.7310
42. Dal Pra A, Lalonde E, Sykes J, Warde F, Ishkanian A, Meng A, Maloff C, Srigley J, Joshua AM, Petrovics G, Van Der Kwast T. TMPRSS2-ERG Status Is Not Prognostic Following Prostate Cancer Radiotherapy: Implications for Fusion Status and DSB RepairTMPRSS2-ERG Status and Radiotherapy. Clinical Cancer Research. 2013 Sep 15;19(18):5202-9. https://doi.org/10.1158/1078-0432.CCR-13-1049
43. Fontugne J, Lee D, Cantaloni C, Barbieri CE, Caffo O, Hanspeter E, Mazzoleni G, Palma PD, Rubin MA, Fellin G, Mosquera JM. Recurrent Prostate Cancer Genomic Alterations Predict Response to Brachytherapy TreatmentProstate Cancer DNA Lesions Predict Brachytherapy Response. Cancer epidemiology, biomarkers & prevention. 2014 Apr 1;23(4):594-600.