Vision Transformers for Diagnostic Classification of Lymphomas: A Matched Comparison with a Convolutional Neural Network
Daniel Rivera, Alexander Banerjee, Rongzhen Zhang, Hanadi El Achi, Amer Wahed, Lauren Ho, and Andy Nguyen*
Department of Pathology and Laboratory Medicine, The University of Texas Health Science Center at Houston, Houston, TX, USA
*Corresponding author:Professor. Andy Nguyen, M.D., Department of Pathology and Laboratory Medicine, The University of Texas Health Science Center at Houston, 6431 Fannin, MSB 2.224, Houston, TX, 77030, USA; E-mail: nghia.d.nguyen@uth.tmc.edu
Received: 19 August 2025; Revised: 04 September 2025; Accepted: 11 September; Published: 16 September 2025
Copyright: © 2025 Rivera D. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Rivera D, Banerjee A, Zhang R, El Achi H, Wahed A, Ho L, et al. (2025) Vision Transformers for Diagnostic Classification of Lymphomas: A Matched Comparison with a Convolutional Neural Network, 21st Century Pathology, Volume 5(1): 160
Abstract
Vision transformers (ViT) have been shown to outperform convolutional neural network (CNN) when pre-trained on sufficient data. ViT has a weaker inductive bias and, therefore, allows for more flexible feature detection. ViT models demonstrate good accuracy on large-scale datasets thanks to their self-supervised learning and multimodal training capabilities. Due to their promising feature detection capabilities, we deployed ViTs for morphological classification of anaplastic large cell lymphoma (ALCL) versus classical Hodgkin lymphoma (cHL). We compared the classification performance of the ViT model with that of our previously designed CNN on the same dataset. Our study presents the first direct comparison of predictive performance between a CNN and a Vision Transformer model, utilizing the same dataset that encompasses ALCL and cHL cases. Our algorithm achieved a diagnostic accuracy of 100% and an F1 score of 1.0 on the independent test set, matching the performance of our previously developed CNN model. The confusion matrix demonstrated perfect classification, with zero false positives or false negatives for both diseases. Our findings suggest that the ViT model can achieve diagnostic performance comparable to that of a CNN, even with small datasets.
Keywords:
Vision transformer; Deep learning; Hodgkin lymphoma; Anaplastic large cell lymphoma
Introduction
Machine learning consists of software that can learn from and make predictions on data-i.e., “gives software the ability to learn without being explicitly programmed” [1]. Numerous machine learning methods have been attempted in the past with varying degrees of success, including decision trees, cluster analysis, support vector machines, random forests, Bayesian analysis, regression analysis, neural networks, and large language models (LLMs). Deep learning (DL) is the most recent and most disruptive method of machine learning, based on neural networks and LLM [2,3]. Currently, many large companies are analyzing large volumes of data for business analysis and decision-making, utilizing deep learning technology (e.g., Google, Microsoft, OpenAI). The application of DL to digital pathology has shown potential that may have an impact on personalized diagnostics and treatment. Breakthroughs in DL occurred in 2006, enabling it to outperform other machine learning models. DL algorithms include two critical features: first, unsupervised learning enables a network to be fed with raw data (with no known outcomes) and to automatically discover the representations needed for detection or classification, and second, supervised learning extracts high-level and complex data representations, thereby avoiding the problems of last-generation networks. DL has significantly benefited from supporting hardware that facilitates parallel computation through numerous graphics processing units (GPU) [4].
Lymphoma is a hematological malignancy, and the current classification of lymphoid malignancies by the World Health Organization is extensive [5]. Worldwide, 280,000 people have been diagnosed with lymphoproliferative neoplasms annually. Lymphoma is typically first suspected based on its growth pattern and the cytologic features of the tumor cells, evaluated by light microscopy of hematoxylin-eosin-stained (H&E) tissue sections. Immunophenotypic features are required for diagnosis, as determined by flow cytometry and/or immunohistochemical stains. Additionally, cytogenetic results, molecular findings, and clinical features are often necessary to confirm the diagnosis or for prognostication. Due to subtle differences in histologic features between various types of lymphomas, it can represent a challenge to pathologists. Applying automated diagnostic tools that utilize whole-slide imaging (WSI) can help pathologists enhance diagnostic precision and efficiency. Earlier attempts to classify histologic images using features such as nuclear shape, size, and texture were largely unsuccessful [6], prompting the adoption of DL algorithms, particularly CNN, which now represent the state-of-the-art in computer vision [7,8].
Convolution in CNN is an operation in image processing that uses filters to modify or detect specific characteristics of an image, such as smoothing, sharpening, intensification, or enhancement. Mathematically, a convolution is done by multiplying the pixel value in the image patch by a filter matrix (kernel) to yield a dot product. By moving the filter across an input image, one obtains the final output as a modified filtered image (Figure 1) [9].
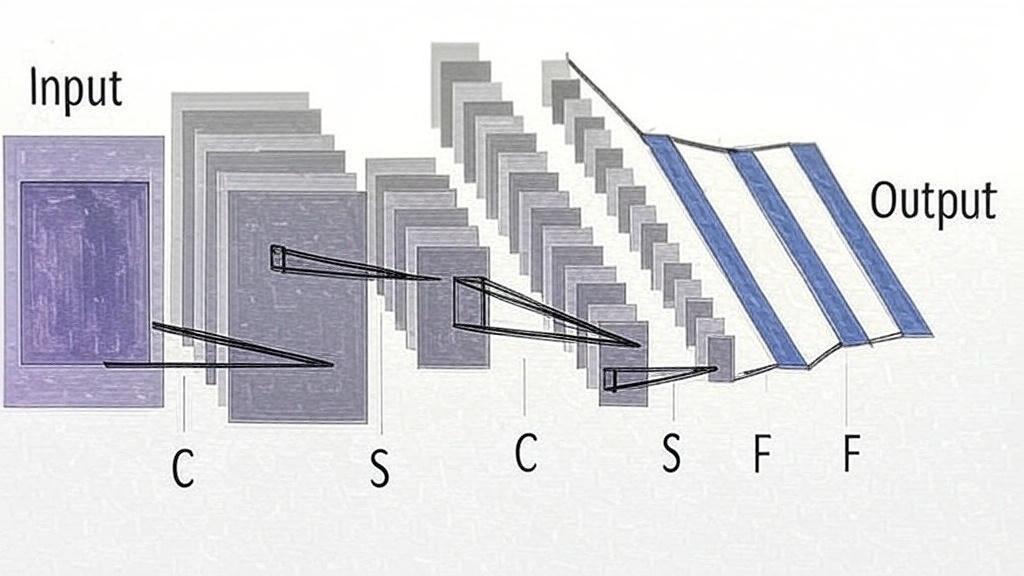
Figure 1: Input: The initial image or data fed into the CNN, represented as a 2D grid (e.g., a pixel array). C (Convolutional layer): A layer that applies convolution operations to extract features like edges or textures, producing feature maps. S (Subsampling/Pooling layer): A layer that reduces the spatial dimensions of feature maps (e.g., via max pooling or average pooling) to simplify computation and improve robustness. F (Fully connected layer): A layer where all neurons are connected to every neuron in the previous layer, used to combine features for the final prediction. Output: The result of the CNN, often a vector of class probabilities or a single prediction.
Recently, ViT has been shown to outperform CNN when pre-trained on enough data [10-12]. In comparison to CNN, ViT exhibits a weaker locality bias, enabling more general feature detection in multimodal data to track attention links between two input tokens. The pixel is the most basic unit of measurement in an image, but calculating every pixel relationship in a normal image would be memory-intensive. ViT takes several steps, which require dividing the entire image into a grid of small image patches, then applying linear projection to embed each patch, with consideration for the position of each image patch in the image; each embedded patch becomes a token, and the resulting sequence of embedded patches is passed to the transformer encoder. The input patches are processed using multi-head attention, and the output is fed into a multilayer perceptron (MLP), which produces the resultant classes, such as tumor types, in a tumor classifier. Furthermore, self-attention in ViT enables each part of the image to relate to other parts, regardless of the distance between them (Figure 2). ViT has been used to build foundation models that are trained on enormous datasets using self-supervised learning, which does not require labeling [13]. They can be fine-tuned for a wide range of downstream tasks using a modest amount of task-specific labeled data for training [10-12].
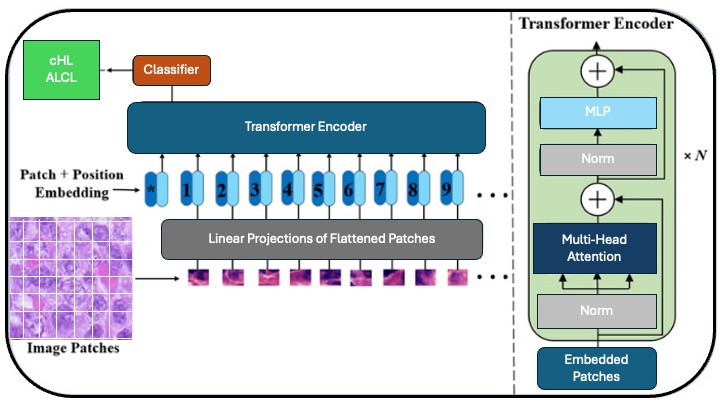
Figure 2: Vision transformer (ViT) for image classification: The Input image is split into smaller patches. Patch+Position Embedding, where patches are flattened, projected into vectors, and combined with positional embeddings. Linear projection converting patches into fixed-size embeddings. The Transformer Encoder utilizes N layers of multi-head attention, fully connected layers (MLP), and layer normalization to process embeddings. Multi-head attention, MLP, and normalization are core components of the Transformer that capture relationships and stabilize the training process. Embedded Patches are a sequence of patch embeddings input to the Transformer. The classifier outputs the final classification (e.g., "Normal" or "Tumor").
The Transformer architecture standard for natural language processing tasks and its applications to computer vision remain limited. In 2021, Dosovitskiy A, et al. deployed a ViT model for image analysis. They demonstrated that a pure transformer applied directly to sequences of image patches can achieve high performance on image classification tasks [12]. We aimed to compare the classification performance of the ViT model with our previously designed CNN on the same dataset [14]. Previous attempts to prove that ViT models can be designed to achieve predictive accuracy comparable to that of CNN, even with small datasets across multiple disciplines in pathology [15-17].
Materials and Methods
We conducted a retrospective collection of cases with newly diagnosed cHL and ALCL, as defined by the current World Health Organization criteria (5th edition), at our institution from 2017 to 2024. We reviewed the morphological characteristics of each case. Next, we obtained WSI of H&E-stained slides from 20 cases (10 cases of ALCL and 10 of cHL), which were scanned using the Philips SG60 scanner (Philips Corporation, Amsterdam, Netherlands) with a scanning capacity for 60 glass slides, which produces high-quality images, full automation for focus, calibration, brightness and contrast settings, with tissue shape detection to outline and scan non-rectangular regions of interest for shorter turnaround times. The total scan time for a 15 × 15 mm benchmark scan area at 40x resolution is less than or equal to 62 seconds. The images were acquired and stored in iSyntax2 format. A Philips Image Management System was used to display the images. From each WSI, 60 image patches of 100 × 100 pixels for feature extraction at 20x magnification, 0.5 μm/pixel using SnagIt software (TechSmith Corp, Okemos, Michigan, USA), yielding a total of 1,200 image patches. Of these, 1080 were used for training, and 120 were used for testing. The cases were divided into two cohorts, with 10 cases for each diagnostic category (Figure 3). For the ViT model, the training set included 540 image patches from ALCL cases and 540 from cHL cases. The test set consisted of 60 image patches from ALCL cases and 60 from cHL cases.
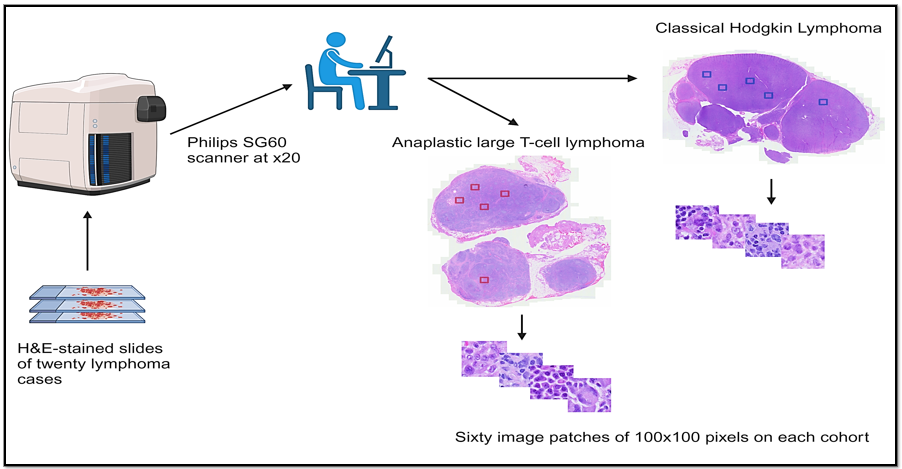
Figure 3: Lymphoma image processing workflow: Twenty lymphoma cases stained with Hematoxylin and Eosin (H&E). Philips SG60 scanner at x20: Scans slides at 20x magnification. Classical Hodgkin lymphoma and anaplastic large T-Cell lymphoma: Two lymphoma types analyzed, shown with marked regions (blue and red squares). Image patches: Sixty 100 × 100 pixel patches were extracted from each lymphoma cohort for detailed analysis.
The algorithm was written in Python using Torch, Torchvision, and Sklearn.metrics libraries (Python Software Foundation, Wilmington, Delaware, USA). Components of the ViT algorithm are shown in Table 1. Parallel processing was performed using an NVIDIA RTX A4000 GPU with 16 GB of memory (NVIDIA Corp, Santa Clara, California, USA) and 6144 CUDA (Compute Unified Device Architecture) cores. Windows 11 Professional, 64-bit was the used operating system (Microsoft Corp, Redmond, Washington, USA.). The Institutional Review Board Ethics Committee approved the protocol for this retrospective study under the principles outlined in the Declaration of Helsinki (Table 1).
Table 1: Components of a vision transformer algorithm.
Steps | Function
Core code for ViT engine | Initialize model parameters, create ViT model, positional encoding (sine & cosine), multi-head attention with query/key/value, attention score, SoftMax, context vector, concatenation, transformer block (ReLU, dropout), define ViT class.
Model & optimizer initialization | Instantiate the Vision transformer, define the loss function, and specify the optimizer (Adam).
Dataset preparation | Load and prepare training and testing datasets.
Model training | Train the ViT model on the training dataset.
Model evaluation | Evaluate the trained model on the testing dataset.
Results visualization | Display all test images alongside known diagnoses and model predictions.
Results
The classification task was binary, distinguishing between ALCL (label 0) and cHL (label 1). The training was conducted for 200 epochs, during which the loss decreased consistently, from 0.256 at Epoch 1 to 0.005 at Epoch 200, indicating effective convergence without overfitting. The ViT training took 3.03 minutes on the NVIDIA RTX A4000, while the CNN took 1.83 minutes. The F1 score remained at a perfect 1.0 throughout 200 epochs, indicating consistent and high-quality performance. Our vision transformer model demonstrated accurate and consistent performance in classifying anaplastic large-cell lymphoma and classical Hodgkin lymphoma. On the independent test set, at Epoch 200, the model achieved a diagnostic accuracy of 100% and an F1 score of 1.0, matching the performance of our previously developed convolutional neural network model, which was also deployed on the same dataset. These results were further visualized in a plot of the F1 score versus the training epoch, which confirmed that the model maintained perfect performance throughout the entire training process (Figure 4).
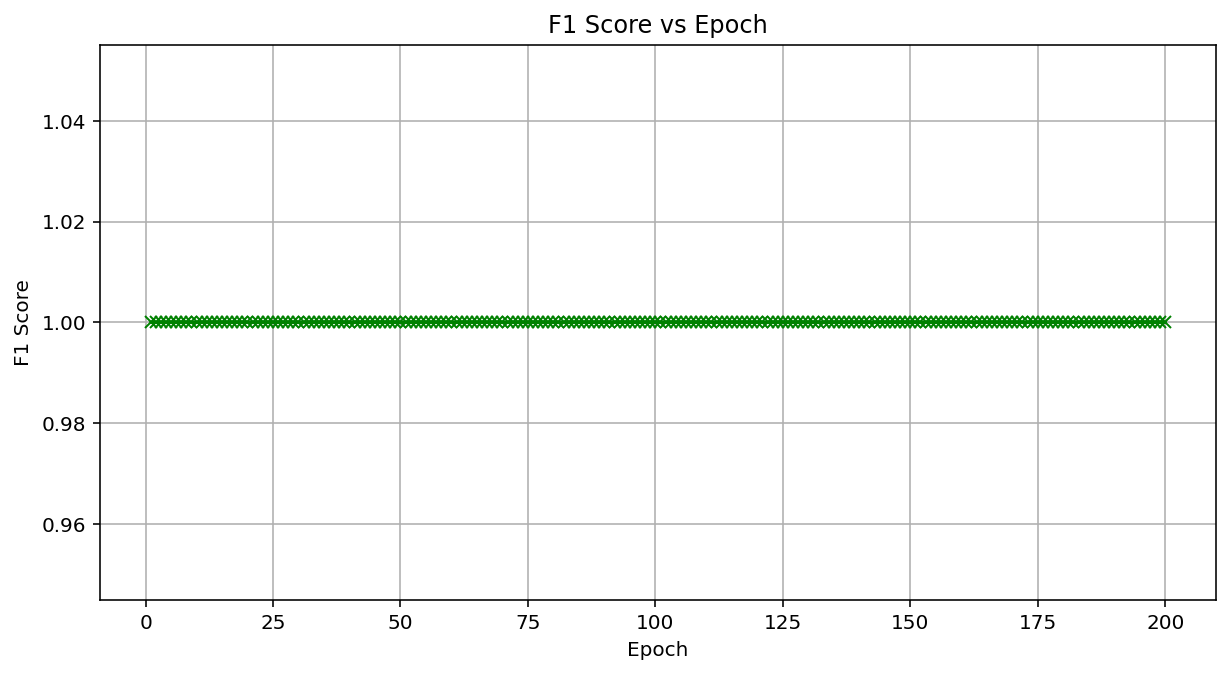
Figure 4: Vision transformer performance by F1 score vs. Epoch for a vision transformer classifying anaplastic large cell lymphoma (0) vs. classical hodgkin lymphoma (1). Model: Built with PyTorch, using multi-head attention, six transformer layers, and 200 epochs. Results: F1 score remains constant at 1.0 across all 200 epochs, indicating perfect classification performance in a dataset including 1080 training and 120 testing images (100 × 100 pixels, 20 × 20 patches).
A confusion matrix was constructed to evaluate the model’s final performance, showing no misclassifications (Figure 5). A production protocol for testing unknown images is established for the ViT-trained model to offer a prediction of diagnosis for new (unknown) images. Figure 6 illustrates a typical screen display for 2 unknown images (Test01.jpg and Test02.jpg). They were both correctly predicted to be ALCL and cHL, respectively.
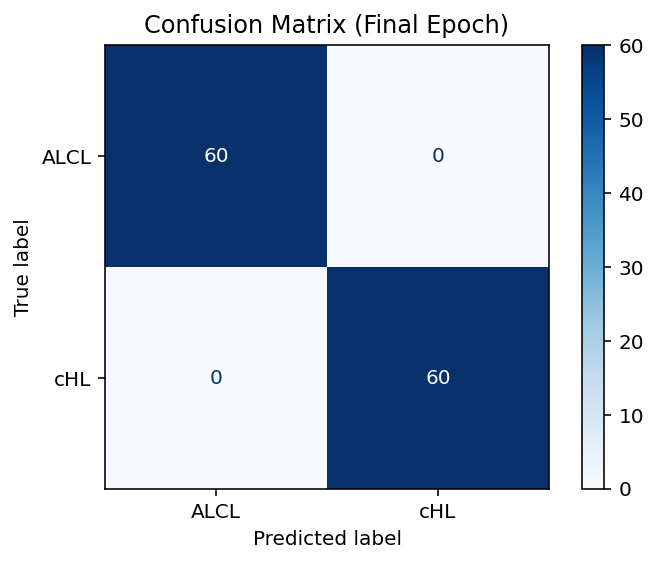
Figure 5: The confusion matrix illustrates the performance of the Vision Transformer at Epoch 200 for ALCL (0) vs. cHL (1) classification. True vs. Predicted Labels: Rows (True: ALCL, CHL); Columns (Predicted: ALCL, cHL). Results: 60/60 ALCL and 60/60 cHL correctly predicted; no misclassifications-performance: 100% accuracy, F1 score of 1.0 on 120 test images.

Figure 6: Screen display of the production model, predicting diagnosis for 2 unknown images.
Discussion
This study presents the first direct comparison between a Vision Transformer and a Convolutional Neural Network for the morphological classification of anaplastic large cell lymphoma and classical Hodgkin lymphoma using the same whole-slide image-derived dataset. Our results demonstrate that the ViT model achieved equivalent performance to the CNN model, an outcome that may be attributed to several key factors. First, all image patches were carefully labelled by hematopathologists to ensure high-quality annotations. Second, the dataset was standardized by sourcing all WSIs from a single laboratory and scanner under consistent imaging conditions. Third, the classification task involved only two well-defined lymphoma types, which likely simplified the discriminative task and enhanced the model’s ability to distinguish between them. Finally, the morphological features of the cells in these lymphoma types tend to be consistent across WSI regions, enabling the model to generalize effectively even from small image patches.
While CNN remains the most widely adopted architecture for medical image classification, due in part to its mature tooling and efficiency on smaller datasets, ViT has shown growing promise in the field of computer vision. Notably, ViT lacks the strong inductive biases of CNN, such as local spatial hierarchies, instead relying on global self-attention mechanisms. This design allows ViT to capture long-range dependencies and potentially generalize better to multimodal or heterogeneous data. However, its performance is often constrained by the algorithm’s reliance on substantially larger datasets to achieve competitive performance, stemming from its data-hungry nature and reduced architectural assumptions.
Transformer-based models are also computationally intensive, particularly when processing large images, such as whole-slide images, where the self-attention mechanism incurs a quadratic computational cost that is directly proportional to the input size. These limitations have historically restricted the application of ViT in digital pathology, where images can span gigapixels. Nonetheless, our study uniquely demonstrates that with proper data curation and task formulation, ViT algorithms can perform comparably to CNN in distinguishing lymphoma types. These findings have proven to hold even on moderately sized datasets, challenging the prevailing view that ViT is restricted to only large-scale pretraining scenarios.
Another limitation of our study is its reliance on supervised learning with fully labelled data, a process that can be time- and labor-intensive. Future directions will include exploring self-supervised pretraining and foundation model development, which may reduce the need for large amounts of labelled data while improving transferability. Additionally, performance should be validated across larger and more diverse datasets, including variations in stain types, scanner brands, and tissue heterogeneity. Cross-institutional validation will be crucial to assessing model robustness in real-world settings.
Conclusion
Our findings support the feasibility and effectiveness of Vision transformers for lymphoma classification tasks in hematopathology. Given their scalability, attention-based modeling, and adaptability to self-supervised learning, the ViT-based architecture represents a promising avenue for future developments in computational pathology. Beyond predictive classification, ViT models hold immense potential as adjuncts in medical education, enabling trainees in pathology to engage with highly accurate computational tools and develop a more nuanced understanding of histologic features relevant to classification. As the field of pathology continues to adopt digital platforms, ViT-guided slide review may supplement both self-directed learning and traditional, instructor-led microscopy sessions, improving overall efficiency and proficiency. In clinical practice, ViT could provide utility as a diagnostic support tool, offering reassurance and reinforcement to pathologists in cases where a diagnosis is uncertain. Furthermore, by highlighting atypical or ambiguous morphologies, this model could serve to identify areas that may warrant additional review. This capability would be particularly valuable in settings with limited access to subspecialty expertise. As these applications materialize, integration into routine diagnostic workflows will require careful validation, optimization, and ongoing clinician oversight to ensure safe and effective use. Ultimately, with continued innovation in reducing their computational demands, ViT may emerge with an increasingly central role in the evolving landscape of digital diagnostics and clinical decision support.
Author Contributions
Daniel Rivera, Alex Banerjee, and Lauren Ho wrote the manuscript. Andy Nguyen mentored, reviewed, and edited the manuscript. All authors approved the manuscript in its final form.
Conflict of Interest
The authors declare that they have no conflicts of interest.
References
- Razzak MI, Naz S, Zaib A. Deep learning for medical image processing: Overview, challenges, and the future. In: Dey N, Ashour A, Borra S. (eds) Classification in BioApps. First ed. Springer International Publishing; 2018:323-350. https://doi.org/10.1007/978-3-319-65981-7_12
- MIT Technol. Rev. 2013. Available at (last accessed on 10/30/18): https://www.technologyreview.com/s/513696/ deep-learning
- LeCun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015 May;521:436–444. https://doi.org/10.1038/nature14539
- Janowczyk A, Madabhushi A. Deep learning for digital pathology image analysis: A comprehensive tutorial with selected use cases. J Pathol Inform. 2016 July;7(1):29. https://doi.org/10.4103/2153-3539.186902
- WHO Classification of Tumours: Haematolymphoid tumours, 5th Edition, Volume 11, 2024. WHO Classification of Tumours Editorial Board. 69008 Lyon, France: International Agency for Research on Cancer (IARC).
- Choras RS. Feature Extraction for CBIR and Biometrics Applications. 7th WSEAS International Conference on Applied Computer Science. 2007 Nov;pp:1-9.
- Marsland S. Machine learning: An algorithmic perspective. Chapman and Hall/CRC, 2011.
- Mitchell TM. Machine learning. McGraw-Hill, New York. 1997;1.
- Roy K, Banik D, Bhattacharjee D, Nasipuri M. Patch-based system for classification of breast histology images using deep learning. Computerized Medical Imaging and Graphics. 2019 Jan;71:90-103.https://doi.org/10.1016/j.compmedimag.2018.11.003
- Waqas A, Bui MM, Glassy EF, El Naqa I, Borkowski P, Borkowski AA, et al. Revolutionizing digital pathology with the power of generative artificial intelligence and foundation models. Lab Invest. 2023 Nov;103(11):100255. https://doi.org/10.1016/j.labinv.2023.100255
- Vaswani A, Shazeer N, Parmar N, Uszkoreit J, Jones L, Gomez AN, et al. Attention Is All You Need. NIPS’17: Proceedings of the 31st Conference on Neural Information Processing Systems. Curran Associates Inc. Red Hook, NY, US. 2017 Dec 4;pp:7104.
- Dosovitskiy A, Beyer L, Kolesnikov A, Weissenborn D, Zhai X, Unterthiner T, et al. An image is worth 16×16 words: Transformers for image recognition at scale. Proceedings of ICLR 2021 Oral. 2021 Jan 12. https://doi.org/10.48550/arXiv.2010.11929
- Vorontsov E, Bozkurt A, Casson A, Shaikovski G, Zelechowski M, Severson K, et al. A foundation model for clinical-grade computational pathology and rare cancers detection. Nature Medicine. 2024 Oct 30;30:2924–2935. https://doi.org/10.1038/s41591-024-03141-0
- Rivera D, Ali K, Zhang R, Mai B, El Achi H, Armstrong J, et al. Deep learning-based morphological classification between classical Hodgkin lymphoma and anaplastic large cell lymphoma: A proof of concept and literature review. 21st Century Pathology. 2024 Nov 18;4(1):159.
- Zhang J, Lu J, Chen B, Pan S, Jin L, Zheng Y, et al. Vision transformer introduces a new vitality to the classification of renal pathology. BMC Nephrology. 2024 Oct 9;25:337. http://dx.doi.org/10.1186/s12882-024-03800-x
- Khedr OS, Wahed ME, Al?Attar AR, Abdel-Rehim EA. The classification of the bladder cancer based on Vision Transformers (ViT). Sci Rep. 2023 Nov 24;13(1):20639. https://doi.org/10.1038/s41598-023-47992-y
- Beyer L, Zhai X, Kolesnikov A. Better plain ViT baselines for ImageNet-1k. Google Research, Brain Team, Zurich. https://github.com/google-research/big_vision.